January 8, 2019
Genomic DNA Preparation by Macherey-Nagel Tissue Spin Columns
Using this kit according to the manufacturer’s protocol is the preferred way for preparing genomic DNA at OTTC when it is intended for ddPCR (i.e. copy-number quantification). Typically, 10 to 90 ng of genomic DNA are used in an 11-25uL PCR reaction.
Genomic DNA Preparation by the “Hot Shot” method
This method was adapted by NIDA-OTTC from the protocol originally described in Biotechniques 29(1), 52-54 (2000). It has been used successfully with ear punches and tail clips of rats. This method is quicker and more cost-effective when compared to the MN Tissue Spin columns, but in our experience, it does not produce templates that are reliable for ddPCR and quantitative analysis. Typically, 1uL of template is sufficient for an 11-25uL PCR reaction.
- Place the biopsy sample in 1.5ml microfuge tube.
- Add 300 microliters of 50mM NaOH
- Incubate tubes at 95C for 60 minutes.
- Vortex tubes on medium power setting for 5 seconds.
- Quick spin the tubes to bring down the condensation.
- Neutralize each sample by adding 30 microliters of 1M Tris-HCl (pH 8).
- Vortex tubes on medium power setting for 5 seconds.
- Quick spin the tubes to bring down the condensation.
Debris (the “undigested” remnant of sample) may remain visible at the bottom of the tube. This is OK, but be sure to take only from the supernatant when setting the PCR reaction.
Use 1uL of this supernatant in a PCR reaction (11uL – 25uL final volume).
General PCR reaction setup:
11.0 uL 2x OneTaq master mix (New England Biolabs)
11.0 uL 2x specific oligos (0.5 uM Forward + 0.5uM Reverse; in water)
3.0 uL genomic DNA
25.0 uL PCR reaction
PCR Program CR2193
| Line | Temp | Time |
| Step 1 | 94°C | HOLD (hot start) |
| Step 2 | 94°C | 2 min |
| Step 3 | 94°C | 30 sec |
| Step 4 | 68°C | 120 sec |
| Step 5 | Go to Step 2 | repeat 34x |
| Step 6 | 68°C | 5 min |
| Step 7 | 12°C | HOLD |
This protocol was updated on 01-08-2019 by CR. Any questions regarding protocol, contact nidatransgenicprojects@mail.nih.gov.
LE-Tg(CX3CR1-Cre-ERT2)3Ottc
Primer Name: CX3CR1 F103289
Primer Sequence (5′ to 3′): TCAGGGTGGCCCATAACCAC
Primer Name: CreERT2 R713
Primer Sequence (5′ to 3′): AGGTAGTTATTCGGATCATCAGCTACAC
These oligos produce a 1017 bp amplicon spanning the 5’ end of the CX3CR1-Cre-ERT2 using with OneTaq polymerase with 68oC annealing.
Primer Name: CX3CR1 F103289
Primer Sequence (5′ to 3′): TCAGGGTGGCCCATAACCAC
Primer Name: CREERT2 R246
Primer Sequence (5′ to 3′): ACCGCGCGCCTGAAGATATAG
These oligos produce a 550 bp amplicon spanning the 5’ end of the CX3CR1-Cre-ERT2 using with OneTaq polymerase with 68oC annealing.
Primer Name: CreERT2 F1206
Primer Sequence (5′ to 3′): CAGTGAAGCTTCGATGATGGGC
Primer Name: CX3CR1 R104157
Primer Sequence (5′ to 3′): GCCAGATTTCCTGCAGGACCT
These oligos produce a 1553 bp amplicon spanning the 3’ end of the CX3CR1-Cre-ERT2 using with OneTaq polymerase with 68oC annealing.
Gel image from CR2193 validation of 5’ (R246) and 3’ junctions.
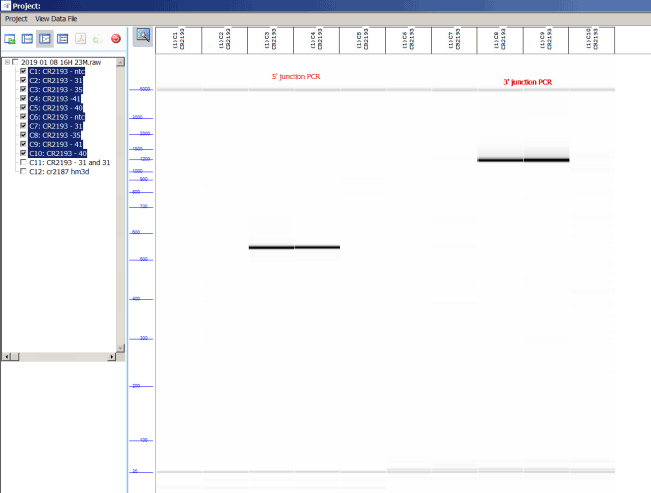
This protocol was updated on 11-28-2017 by CR. Any questions regarding protocol, contact nidatransgenicprojects@mail.nih.gov.
